A new global dataset of 239 human-infecting RNA viruses shows how animal hosts, vector transmission, surveillance gaps, and viral traits shape the pathway from dispersal to epidemic threat.
Study: Complete catalog of human infectious RNA viruses. Image credit: Andrzej Rostek / Shutterstock
A recent review published in the magazine Scientific Data presents an updated global catalog that brings the number of ribonucleic acid (RNA) viruses known to infect humans in 239 species, 25 more than in 2018, offering new insight into emergence and spread.
Rather than appearing randomly, most viruses cluster in a few families, are associated with non-human hosts, particularly mammals, and are identified at varying rates over time as taxonomy, reporting, surveillance, and sequencing technologies evolve.
While spillover to humans is common worldwide, only a minority of species reach epidemic or endemic levels in humans, highlighting a critical bottleneck between exposure and epidemic spread.
RNA Viruses remain a growing threat to global health today, driving diseases such as measles, influenza and AIDS caused by HIV and causing new outbreaks. Recent events involving Oropouche virus and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) highlight the epidemic potential of these viruses. However, the viral landscape continues to change rapidly.
Researchers identify new species that infect humans almost every year, revise classifications, and expand genomic and ecological data. As evidence accumulates about transmission, host range, and dissemination, the need for an updated inventory becomes critical for tracking known and predicting future risks.
Human RNA virus catalog methods
In this paper, the researchers developed an updated, expanded dataset of it RNA viruses known to infect humans, capturing current knowledge through December 2024.
Based on older catalogs from 2001 and 2018, they conducted systematic literature searches every 1–3 years using databases such as Web of Science, PubMed, Scopus and Google Scholar, supplemented by sources including the World Health Organization (WHERE), Centers for Disease Control and Prevention (CDC), ProMed and genome archives from the National Center for Biotechnology Information (NCBI).
The dataset included only peer-reviewed primary reports that provided reliable evidence RNA viruses recognized by the International Committee on the Taxonomy of Viruses (ICTV) infects humans under natural or actual conditions, excluding intentional experimental inoculation or in vitro evidence.
The team resolved ambiguities through independent reviews and consensus, and in some cases inferred features missing from closely related viruses. They compiled species-level data incorporating information on known subtypes and linked each virus to its first reported human case, genome sequence and geographic origin.
The researchers recorded key characteristics, including transmissibility, host range and routes of transmission, using standardized criteria. They classified transmissibility into Levels 2, 3 and 4, ranging from zoonotic infections without human spread to viruses capable of epidemic or endemic spread in humans.
Finally, the team mapped discovery dates and locations, allowing for temporal and spatial analyzes of the virus’s emergence. By integrating genomic, ecological, and epidemiological data into a single framework, the updated dataset provides a robust foundation for the study of viral diversity, evolution, and public health risk.
The annual number of new and (currently) ICTV-recognized, human-infectious RNA virus species is shown in Figure 2a. Figure 2b illustrates the accumulation of species over time, as well as the accumulation of genera and families containing one or more human-infecting RNA virus species. The first human RNA virus – the yellow fever virus – was reported in 1901. The number of species increased slowly until the mid-1950s and somewhat more rapidly thereafter. By the end of the 20th century, 178 species had been identified, and in the 21st century so far another 61 have been added. By decade, the 1960s contributed the largest number of new species (42). Next was the 2000s (31), but the percentage dropped again in the 2010s.
Global RNA virus discovery standards
The updated data set includes 239 RNA viruses known to infect humans, as classified by ICTV. Compared to 2018, this reflects 25 additional species identified through new discoveries and taxonomic updates.
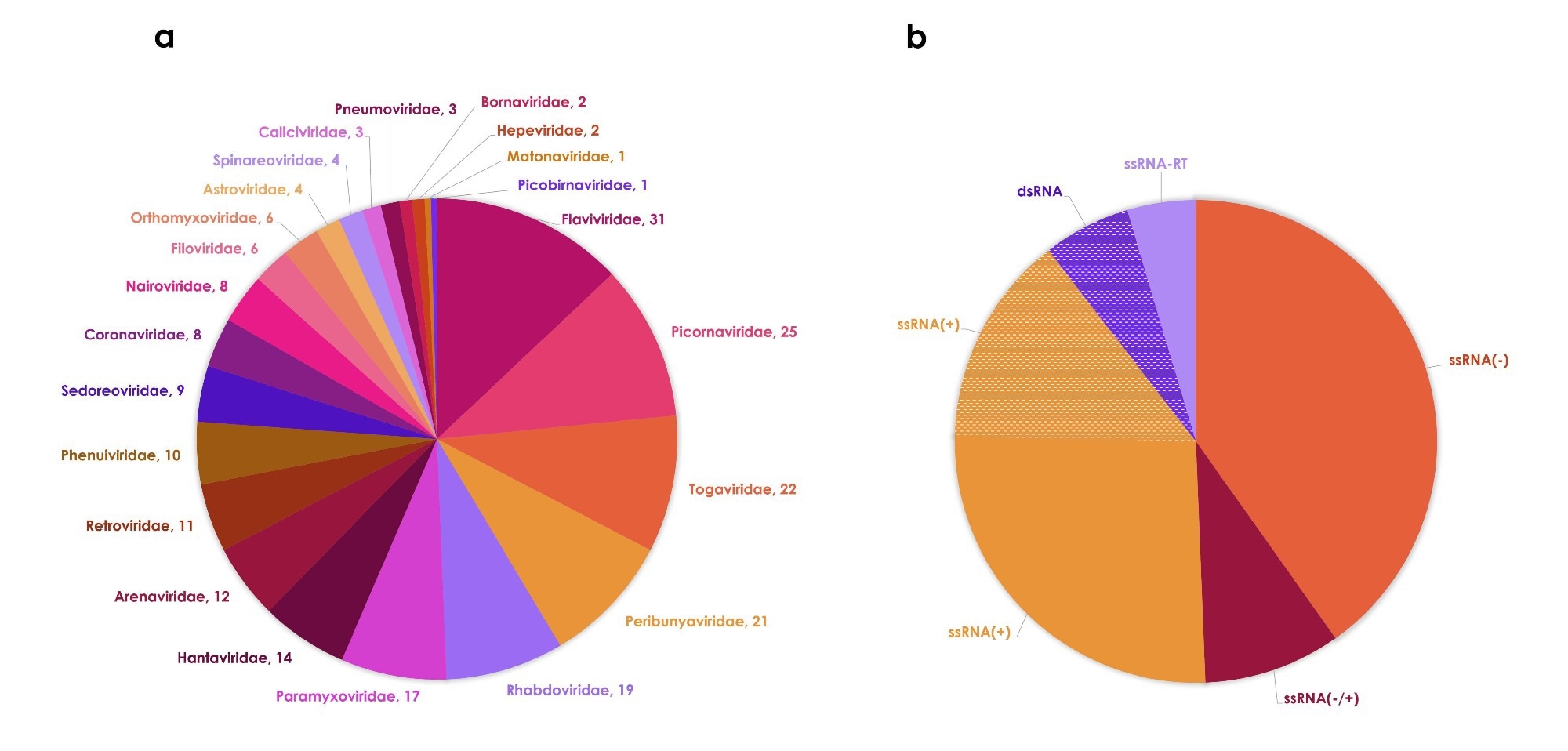
These species span 61 genera and 23 families, although diversity remains concentrated in a few families and most viruses share common genomic features, particularly single-stranded RNA genomes.
Over time, discoveries have increased since the mid-20th century, although the authors note that a formal analysis is needed to determine whether discovery rates are increasing or decreasing overall.
After minimal growth in the early 20th century, recognition rates rose sharply from the mid-1900s, with notable peaks in the 1960s and early 2000s. However, most newly identified species expand existing genera and families rather than introducing entirely new taxonomic groups.
Geographically, the first reported human cases have occurred on all inhabited continents, with clusters in areas with stronger surveillance systems. This pattern highlights both the global nature of viral diffusion and the effect of detectability on discovery.
Spread, Transmission and Epidemic Potential
Ecologically, the majority of viruses (62%) are strictly zoonotic (Level 2) and do not sustain human-to-human transmission. Only 60 species reach level 4, meaning they are endemic to humans or can spread epidemically, and many of these still maintain animal reservoirs.
Most viruses are associated with nonhuman mammalian hosts, reinforcing their central role in emergence. Transmission routes are varied, but vector-borne spread, mainly through mosquitoes and ticks, predominates, followed by inhalation and direct contact.
In particular, the transmission routes of a subset of viruses remain uncertain, reflecting persistent knowledge gaps. Together, these findings clearly highlight a landscape defined by repeated documented spills, expanding discovery, and limited adaptation to sustained human transmission.
RNA virus surveillance and risk prediction
These findings suggest a more targeted and proactive approach to emerging viral threats. Rather than searching broadly for entirely new pathogens, future efforts could use this data set to examine high-risk virus families, mammalian reservoirs, and areas with limited surveillance, where undetected spillover is most likely to occur.
Expanding genomic sequencing, metagenomics and real-time surveillance will be critical to closing persistent knowledge gaps, particularly around transmission routes and host range.
At the same time, the dataset provides a valuable basis for modeling trends in the discovery and identification of traits associated with outbreak potential. As it continues to evolve, it can help improve risk prediction and guide early warning systems.
Ultimately, the challenge is not just to discover new viruses, but to understand which ones are most likely to adapt, spread and become the next global health threat.
